
Top ▲

| Quaternary Structure: Complexes |
| DNA gyrase |
Gene and Protein Information  |
||||||
| Species | TM | AA | Chromosomal Location | Gene Symbol | Gene Name | Reference |
| Escherichia coli | - | 875 | gyrA | DNA gyrase subunit A | ||
Database Links  |
|
| Alphafold | P0AES4 (E.coli) |
| ChEMBL Target | CHEMBL1858 (E.coli) |
| Entrez Gene | 946614 (E.coli) |
| RefSeq Protein | NP_416734 (E.coli) |
| UniProtKB | P0AES4 (E.coli) |
Selected 3D Structures  |
|||||||||||
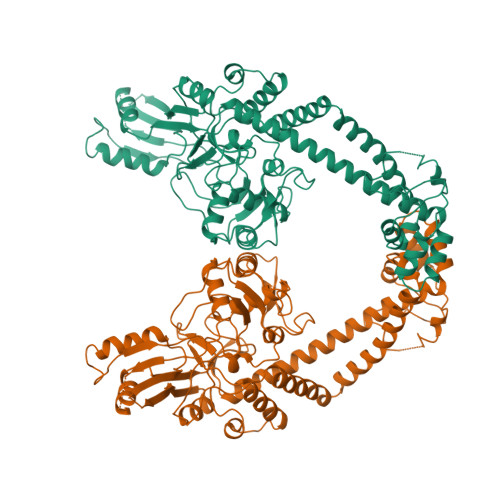
|
|
||||||||||
Download all structure-activity data for this target as a CSV file 
| Inhibitors | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Key to terms and symbols | View all chemical structures | Click column headers to sort | |||||||||||||||||||||||||||||||||||||||||||||||||
|
|||||||||||||||||||||||||||||||||||||||||||||||||||
| General Comments |
| DNA gyrase subunit A is comprised of an N-terminal domain (59-64 kDa) involved in DNA cleavage and ligation, and a C-terminal domain (33 kDa) involved in DNA-protein interactions [3]. |
1. Barnard FM, Maxwell A. (2001) Interaction between DNA gyrase and quinolones: effects of alanine mutations at GyrA subunit residues Ser(83) and Asp(87). Antimicrob Agents Chemother, 45 (7): 1994-2000. [PMID:11408214]
2. Morais Cabral JH, Jackson AP, Smith CV, Shikotra N, Maxwell A, Liddington RC. (1997) Crystal structure of the breakage-reunion domain of DNA gyrase. Nature, 388 (6645): 903-6. [PMID:9278055]
3. Reece RJ, Maxwell A. (1989) Tryptic fragments of the Escherichia coli DNA gyrase A protein. J Biol Chem, 264 (33): 19648-53. [PMID:2555327]
Bacterial protein targets: DNA gyrase subunit A. Last modified on 28/04/2023. Accessed on 10/05/2024. IUPHAR/BPS Guide to PHARMACOLOGY, https://www.guidetopharmacology.org/GRAC/ObjectDisplayForward?objectId=3217.