
GtoPdb is requesting financial support from commercial users. Please see our sustainability page for more information.
Adenosine receptors: Introduction
General
Adenosine is ubiquitously present in the extracellular medium in all tissues and acts as a local modulator by activating four well-defined G protein-coupled receptors [16-18,45,47]. These receptors are denoted adenosine receptors, but may also be called P1 receptors to distinguish them from the receptors for nucleotides, which belong either to the family of transmitter-gated ion channels (P2X receptors) or the family of G protein-coupled receptors (P2Y receptors) [18].
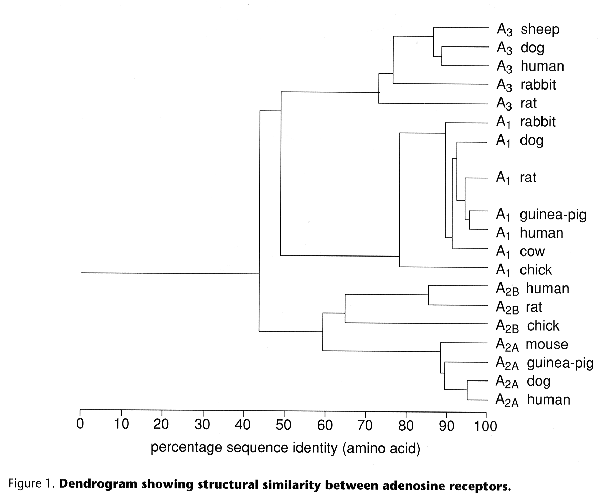
The four adenosine receptors, called A1, A2A, A2B and A3, have been cloned from numerous mammalian species, including man. As seen in Figure 1 there is extensive sequence similarity between species for the A1, and A2A receptors, whereas A2B and A3 receptors are more variable.
Binding sites
There are reports of binding sites with a pharmacology that is not typical for any of the cloned subtypes [12-13,27-28,39]. These data could indicate that there are as yet undiscovered adenosine receptors. However, no evidence for additional adenosine receptors has evolved from the extensive cloning efforts of several laboratories. The A2A receptor has been crystallized and its structure has been determined in complex with several agonist and antagonist ligands [11,15,24-25,34,59]. This structure has been used effectively for in silico screening of chemically diverse libraries to identify antagonists that have novel chemotypes [10,29,33,35,53,58] and also for the construction of reliable homology models of other adenosine receptor subtypes [6,32,49].
Ligands
Over the past decades highly selective agonists and antagonists for adenosine receptors have been developed. Some examples of such compounds are listed in Table 1. For structures and additional information on compounds the reader is referred to recent papers and reviews [2-3,14,26,38,46,51]. For reasons of space only a few representative compounds are given in the following data tables. Combined site-directed mutagenesis and homology-based modelling have identified amino acids in the third and seventh transmembrane domains (TM) as being critically important in agonist binding [30,48]. Much of the previous molecular modelling, as informed by studies of mutagenesis and structure activity relationships (SAR), is an agreement with data from the X-ray crystallographic determination of the A2A adenosine receptor structure.
There is good evidence for the adenosine receptors (as for other 7TM receptors) that agonist potency is strongly dependent on receptor number [1,19,50] and on the assay used. Hence, data for both absolute and relative potencies for agonists must be used cautiously. In particular, the large differences in the number of receptors of different types in vivo could lead to much higher or lower selectivity than suggested by in vitro data.
There is solid evidence that there are partial agonists for all four adenosine receptor subtypes [5,40,55-57]. As for other G protein-coupled receptors, some of the antagonists have been proven to be inverse agonists [4,22,37]

Allosteric site
Besides the traditional agonist recognition site there is evidence, in the case of the adenosine A1 receptor, for an allosteric site that regulates agonist binding and biological effects [8-9,36,41]. The positive allosteric modulator (PAM) PD81723, a thienopyridine derivative, may act by stabilising the receptor-G protein interaction [7], and the same may apply for analogues that are superior in potency and/or selectivity [54]. It is possible that one reason why such allosteric enhancement of A1 receptor agonist binding can be found is related to the fact that these receptors are coupled in a somewhat unusual way to G proteins that may involve some other protein factors [43]. PAMs of the adenosine A3 receptor have also been reported [20,23,31]. Recently bitopic ligands, most likely occupuying orthosteric and allosteric binding sites in the adenosine A1 receptor simultaneously, have been introduced [44,52].
Adenosine in inflammation
All immune cells express receptors for adenosine [21]. It is suggested that adenosine exhibits pro-inflammatory effects through its A1 receptor, and anti-inflammatory effects through A2A receptor. It is noteworthy that A1 receptor expression peaks during leukocyte recruitment, and this drives increased expression of A2A receptor which facilitates resolution of inflammation [42].
References
1. Arslan G, Kull B, Fredholm BB. (1999) Signaling via A2A adenosine receptor in four PC12 cell clones. Naunyn Schmiedebergs Arch Pharmacol, 359 (1): 28-32. [PMID:9933147]
2. Baraldi PG, Cacciari B, Romagnoli R, Merighi S, Varani K, Borea PA, Spalluto G. (2000) A(3) adenosine receptor ligands: history and perspectives. Med Res Rev, 20 (2): 103-28. [PMID:10723024]
3. Baraldi PG, Cacciari B, Spalluto G, Ji XD, Olah ME, Stiles G, Dionisotti S, Zocchi C, Ongini E, Jacobson KA. (1996) Novel N6-(substituted-phenylcarbamoyl)adenosine-5'-uronamides as potent agonists for A3 adenosine receptors. J Med Chem, 39 (3): 802-6. [PMID:8576924]
4. Bennett KA, Tehan B, Lebon G, Tate CG, Weir M, Marshall FH, Langmead CJ. (2013) Pharmacology and structure of isolated conformations of the adenosine A₂A receptor define ligand efficacy. Mol Pharmacol, 83 (5): 949-58. [PMID:23429888]
5. Beukers MW, Chang LC, von Frijtag Drabbe Künzel JK, Mulder-Krieger T, Spanjersberg RF, Brussee J, IJzerman AP. (2004) New, non-adenosine, high-potency agonists for the human adenosine A2B receptor with an improved selectivity profile compared to the reference agonist N-ethylcarboxamidoadenosine. J Med Chem, 47 (15): 3707-9. [PMID:15239649]
6. Beuming T, Sherman W. (2012) Current assessment of docking into GPCR crystal structures and homology models: successes, challenges, and guidelines. J Chem Inf Model, 52 (12): 3263-77. [PMID:23121495]
7. Bhattacharya S, Linden J. (1995) The allosteric enhancer, PD 81,723, stabilizes human A1 adenosine receptor coupling to G proteins. Biochim Biophys Acta, 1265 (1): 15-21. [PMID:7857979]
8. Bruns RF, Fergus JH. (1990) Allosteric enhancement of adenosine A1 receptor binding and function by 2-amino-3-benzoylthiophenes. Mol Pharmacol, 38 (6): 939-49. [PMID:2174510]
9. Bruns RF, Fergus JH, Coughenour LL, Courtland GG, Pugsley TA, Dodd JH, Tinney FJ. (1990) Structure-activity relationships for enhancement of adenosine A1 receptor binding by 2-amino-3-benzoylthiophenes. Mol Pharmacol, 38 (6): 950-8. [PMID:2250667]
10. Carlsson J, Yoo L, Gao ZG, Irwin JJ, Shoichet BK, Jacobson KA. (2010) Structure-based discovery of A2A adenosine receptor ligands. J Med Chem, 53 (9): 3748-55. [PMID:20405927]
11. Congreve M, Andrews SP, Doré AS, Hollenstein K, Hurrell E, Langmead CJ, Mason JS, Ng IW, Tehan B, Zhukov A et al.. (2012) Discovery of 1,2,4-triazine derivatives as adenosine A(2A) antagonists using structure based drug design. J Med Chem, 55 (5): 1898-903. [PMID:22220592]
12. Cornfield LJ, Hu S, Hurt SD, Sills MA. (1992) [3H]2-phenylaminoadenosine ([3H]CV 1808) labels a novel adenosine receptor in rat brain. J Pharmacol Exp Ther, 263 (2): 552-61. [PMID:1331404]
13. Cunha RA, Johansson B, Constantino MD, Sebastião AM, Fredholm BB. (1996) Evidence for high-affinity binding sites for the adenosine A2A receptor agonist [3H] CGS 21680 in the rat hippocampus and cerebral cortex that are different from striatal A2A receptors. Naunyn Schmiedebergs Arch Pharmacol, 353 (3): 261-71. [PMID:8692280]
14. Daly JW, Jacobson KA. (1995) Adenosine receptors: selective agonists and antagonists. In Adenosine and adenine nucleotides: from molecular biology to integrative physiology. Edited by Belardinelli L, Pelleg A (Kluwer Academic Publishers) 157-166. [ISBN:0792331907]
15. Doré AS, Robertson N, Errey JC, Ng I, Hollenstein K, Tehan B, Hurrell E, Bennett K, Congreve M, Magnani F et al.. (2011) Structure of the adenosine A(2A) receptor in complex with ZM241385 and the xanthines XAC and caffeine. Structure, 19 (9): 1283-93. [PMID:21885291]
16. Fredholm BB. (1995) Astra Award Lecture. Adenosine, adenosine receptors and the actions of caffeine. Pharmacol Toxicol, 76 (2): 93-101. [PMID:7746802]
17. Fredholm BB, Abbracchio MP, Burnstock G, Daly JW, Harden TK, Jacobson KA, Leff P, Williams M. (1994) Nomenclature and classification of purinoceptors. Pharmacol Rev, 46 (2): 143-56. [PMID:7938164]
18. Fredholm BB, Abbracchio MP, Burnstock G, Dubyak GR, Harden TK, Jacobson KA, Schwabe U, Williams M. (1997) Towards a revised nomenclature for P1 and P2 receptors. Trends Pharmacol Sci, 18 (3): 79-82. [PMID:9133776]
19. Gerwins P, Fredholm BB. (1991) Glucocorticoid receptor activation leads to up-regulation of adenosine A1 receptors and down-regulation of adenosine A2 responses in DDT1 MF-2 smooth muscle cells. Mol Pharmacol, 40 (2): 149-55. [PMID:1652051]
20. Göblyös A, Gao ZG, Brussee J, Connestari R, Santiago SN, Ye K, Ijzerman AP, Jacobson KA. (2006) Structure-activity relationships of new 1H-imidazo[4,5-c]quinolin-4-amine derivatives as allosteric enhancers of the A3 adenosine receptor. J Med Chem, 49 (11): 3354-61. [PMID:16722654]
21. Haskó G, Linden J, Cronstein B, Pacher P. (2008) Adenosine receptors: therapeutic aspects for inflammatory and immune diseases. Nat Rev Drug Discov, 7 (9): 759-70. [PMID:18758473]
22. He W, Wilder T, Cronstein BN. (2013) Rolofylline, an adenosine A1 receptor antagonist, inhibits osteoclast differentiation as an inverse agonist. Br J Pharmacol, 170 (6): 1167-76. [PMID:23962057]
23. Heitman LH, Göblyös A, Zweemer AM, Bakker R, Mulder-Krieger T, van Veldhoven JP, de Vries H, Brussee J, Ijzerman AP. (2009) A series of 2,4-disubstituted quinolines as a new class of allosteric enhancers of the adenosine A3 receptor. J Med Chem, 52 (4): 926-31. [PMID:19161279]
24. Hino T, Arakawa T, Iwanari H, Yurugi-Kobayashi T, Ikeda-Suno C, Nakada-Nakura Y, Kusano-Arai O, Weyand S, Shimamura T, Nomura N et al.. (2012) G-protein-coupled receptor inactivation by an allosteric inverse-agonist antibody. Nature, 482 (7384): 237-40. [PMID:22286059]
25. Jaakola VP, Griffith MT, Hanson MA, Cherezov V, Chien EY, Lane JR, Ijzerman AP, Stevens RC. (2008) The 2.6 angstrom crystal structure of a human A2A adenosine receptor bound to an antagonist. Science, 322 (5905): 1211-7. [PMID:18832607]
26. Jacobson KA, Kim HO, Siddiqi SM, Olah ME, Stiles GL, von Lubitz DKJE. (1995) A3-adenosine receptors: design of selective ligands and therapeutic prospects. Drugs Future, 20: 689-699.
27. Johansson B, Fredholm BB. (1995) Further characterization of the binding of the adenosine receptor agonist [3H]CGS 21680 to rat brain using autoradiography. Neuropharmacology, 34 (4): 393-403. [PMID:7566470]
28. Johansson B, Georgiev V, Parkinson FE, Fredholm BB. (1993) The binding of the adenosine A2 receptor selective agonist [3H]CGS 21680 to rat cortex differs from its binding to rat striatum. Eur J Pharmacol, 247 (2): 103-10. [PMID:8281998]
29. Katritch V, Jaakola VP, Lane JR, Lin J, Ijzerman AP, Yeager M, Kufareva I, Stevens RC, Abagyan R. (2010) Structure-based discovery of novel chemotypes for adenosine A(2A) receptor antagonists. J Med Chem, 53 (4): 1799-809. [PMID:20095623]
30. Kim J, Wess J, van Rhee AM, Schöneberg T, Jacobson KA. (1995) Site-directed mutagenesis identifies residues involved in ligand recognition in the human A2a adenosine receptor. J Biol Chem, 270 (23): 13987-97. [PMID:7775460]
31. Kim Y, de Castro S, Gao ZG, Ijzerman AP, Jacobson KA. (2009) Novel 2- and 4-substituted 1H-imidazo[4,5-c]quinolin-4-amine derivatives as allosteric modulators of the A3 adenosine receptor. J Med Chem, 52 (7): 2098-108. [PMID:19284749]
32. Kolb P, Phan K, Gao ZG, Marko AC, Sali A, Jacobson KA. (2012) Limits of ligand selectivity from docking to models: in silico screening for A(1) adenosine receptor antagonists. PLoS ONE, 7 (11): e49910. [PMID:23185482]
33. Langmead CJ, Andrews SP, Congreve M, Errey JC, Hurrell E, Marshall FH, Mason JS, Richardson CM, Robertson N, Zhukov A et al.. (2012) Identification of novel adenosine A(2A) receptor antagonists by virtual screening. J Med Chem, 55 (5): 1904-9. [PMID:22250781]
34. Lebon G, Warne T, Edwards PC, Bennett K, Langmead CJ, Leslie AG, Tate CG. (2011) Agonist-bound adenosine A2A receptor structures reveal common features of GPCR activation. Nature, 474 (7352): 521-5. [PMID:21593763]
35. Lenselink EB, Beuming T, Sherman W, van Vlijmen HW, IJzerman AP. (2014) Selecting an optimal number of binding site waters to improve virtual screening enrichments against the adenosine A2A receptor. J Chem Inf Model, 54 (6): 1737-46. [PMID:24835542]
36. Leung E, Walsh LK, Flippin LA, Kim EJ, Lazar DA, Seran CS, Wong EH, Eglen RM. (1995) Enhancement of adenosine A1 receptor functions by benzoylthiophenes in guinea pig tissues in vitro. Naunyn Schmiedebergs Arch Pharmacol, 352 (2): 206-12. [PMID:7477445]
37. Li Q, Ye K, Blad CC, den Dulk H, Brouwer J, Ijzerman AP, Beukers MW. (2007) ZM241385, DPCPX, MRS1706 are inverse agonists with different relative intrinsic efficacies on constitutively active mutants of the human adenosine A2B receptor. J Pharmacol Exp Ther, 320 (2): 637-45. [PMID:17077318]
38. Linden J, Thai T, Figler H, Jin X, Robeva AS. (1999) Characterization of human A(2B) adenosine receptors: radioligand binding, western blotting, and coupling to G(q) in human embryonic kidney 293 cells and HMC-1 mast cells. Mol Pharmacol, 56 (4): 705-13. [PMID:10496952]
39. Lindström K, Ongini E, Fredholm BB. (1996) The selective adenosine A2A receptor antagonist SCH 58261 discriminates between two different binding sites for [3H]-CGS 21680 in the rat brain. Naunyn Schmiedebergs Arch Pharmacol, 354 (4): 539-41. [PMID:8897460]
40. Mathôt RA, Van der Wenden EM, Soudijn W, IJzerman AP, Danhof M. (1995) Deoxyribose analogues of N6-cyclopentyladenosine (CPA): partial agonists at the adenosine A1 receptor in vivo. Br J Pharmacol, 116 (3): 1957-64. [PMID:8640332]
41. Mudumbi RV, Montamat SC, Bruns RF, Vestal RE. (1993) Cardiac functional responses to adenosine by PD 81,723, an allosteric enhancer of the adenosine A1 receptor. Am J Physiol, 264 (3 Pt 2): H1017-22. [PMID:8456969]
42. Nakav S, Chaimovitz C, Sufaro Y, Lewis EC, Shaked G, Czeiger D, Zlotnik M, Douvdevani A. (2008) Anti-inflammatory preconditioning by agonists of adenosine A1 receptor. PLoS ONE, 3 (5): e2107. [PMID:18461129]
43. Nanoff C, Mitterauer T, Roka F, Hohenegger M, Freissmuth M. (1995) Species differences in A1 adenosine receptor/G protein coupling: identification of a membrane protein that stabilizes the association of the receptor/G protein complex. Mol Pharmacol, 48 (5): 806-17. [PMID:7476910]
44. Narlawar R, Lane JR, Doddareddy M, Lin J, Brussee J, Ijzerman AP. (2010) Hybrid ortho/allosteric ligands for the adenosine A(1) receptor. J Med Chem, 53 (8): 3028-37. [PMID:20345101]
45. Olah ME, Stiles GL. (1995) Adenosine receptor subtypes: characterization and therapeutic regulation. Annu Rev Pharmacol Toxicol, 35: 581-606. [PMID:7598508]
46. Ongini E, Fredholm BB. (1996) Pharmacology of adenosine A2A receptors. Trends Pharmacol Sci, 17 (10): 364-72. [PMID:8979771]
47. Palmer TM, Stiles GL. (1995) Adenosine receptors. Neuropharmacology, 34 (7): 683-94. [PMID:8532135]
48. Rivkees SA, Barbhaiya H, IJzerman AP. (1999) Identification of the adenine binding site of the human A1 adenosine receptor. J Biol Chem, 274 (6): 3617-21. [PMID:9920910]
49. Sherbiny FF, Schiedel AC, Maass A, Müller CE. (2009) Homology modelling of the human adenosine A2B receptor based on X-ray structures of bovine rhodopsin, the beta2-adrenergic receptor and the human adenosine A2A receptor. J Comput Aided Mol Des, 23 (11): 807-28. [PMID:19757091]
50. Shryock JC, Snowdy S, Baraldi PG, Cacciari B, Spalluto G, Monopoli A, Ongini E, Baker SP, Belardinelli L. (1998) A2A-adenosine receptor reserve for coronary vasodilation. Circulation, 98 (7): 711-8. [PMID:9715864]
51. Svenningsson P, Le Moine C, Fisone G, Fredholm BB. (1999) Distribution, biochemistry and function of striatal adenosine A2A receptors. Prog Neurobiol, 59 (4): 355-96. [PMID:10501634]
52. Valant C, May LT, Aurelio L, Chuo CH, White PJ, Baltos JA, Sexton PM, Scammells PJ, Christopoulos A. (2014) Separation of on-target efficacy from adverse effects through rational design of a bitopic adenosine receptor agonist. Proc Natl Acad Sci USA, 111 (12): 4614-9. [PMID:24619092]
53. van der Horst E, van der Pijl R, Mulder-Krieger T, Bender A, Ijzerman AP. (2011) Substructure-based virtual screening for adenosine A2A receptor ligands. ChemMedChem, 6 (12): 2302-11. [PMID:22021213]
54. van der Klein PA, Kourounakis AP, IJzerman AP. (1999) Allosteric modulation of the adenosine A(1) receptor. Synthesis and biological evaluation of novel 2-amino-3-benzoylthiophenes as allosteric enhancers of agonist binding. J Med Chem, 42 (18): 3629-35. [PMID:10479294]
55. van der Wenden EM, Carnielli M, Roelen HC, Lorenzen A, von Frijtag Drabbe Künzel JK, IJzerman AP. (1998) 5'-substituted adenosine analogs as new high-affinity partial agonists for the adenosine A1 receptor. J Med Chem, 41 (1): 102-8. [PMID:9438026]
56. van Tilburg EW, Gremmen M, von Frijtag Drabbe Künzel J, de Groote M, IJzerman AP. (2003) 2,8-Disubstituted adenosine derivatives as partial agonists for the adenosine A2A receptor. Bioorg Med Chem, 11 (10): 2183-92. [PMID:12713828]
57. van Tilburg EW, von Frijtag Drabbe Kunzel J, de Groote M, IJzerman AP. (2002) 2,5'-Disubstituted adenosine derivatives: evaluation of selectivity and efficacy for the adenosine A(1), A(2A), and A(3) receptor. J Med Chem, 45 (2): 420-9. [PMID:11784146]
58. van Westen GJ, van den Hoven OO, van der Pijl R, Mulder-Krieger T, de Vries H, Wegner JK, Ijzerman AP, van Vlijmen HW, Bender A. (2012) Identifying novel adenosine receptor ligands by simultaneous proteochemometric modeling of rat and human bioactivity data. J Med Chem, 55 (16): 7010-20. [PMID:22827545]
59. Xu F, Wu H, Katritch V, Han GW, Jacobson KA, Gao ZG, Cherezov V, Stevens RC. (2011) Structure of an agonist-bound human A2A adenosine receptor. Science, 332 (6027): 322-7. [PMID:21393508]







