
Top ▲

GtoPdb is requesting financial support from commercial users. Please see our sustainability page for more information.
Database Links  |
|
| ChEMBL Target | CHEMBL4523582 (SARS-CoV-2), CHEMBL5118 (SARS-CoV) |
| UniProtKB | P0DTD1 (SARS-CoV-2), P0C6X7 (SARS-CoV) |
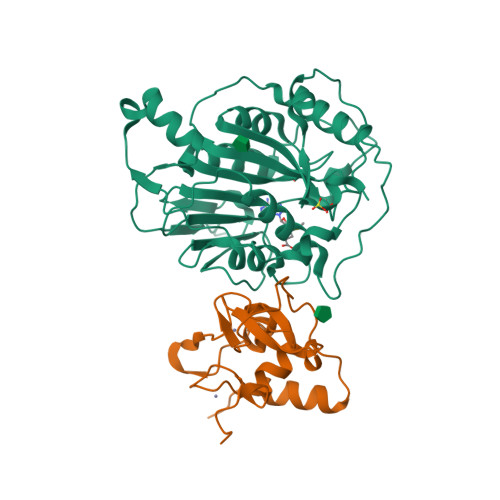
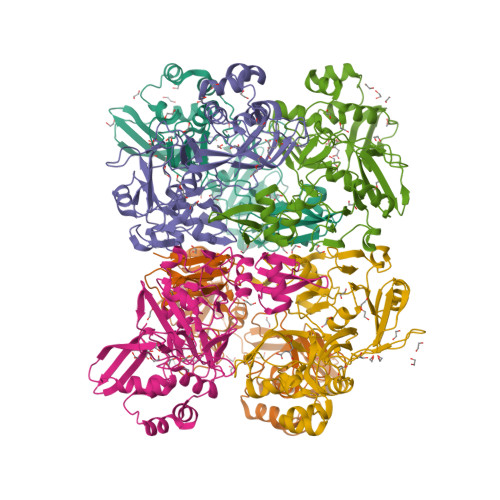
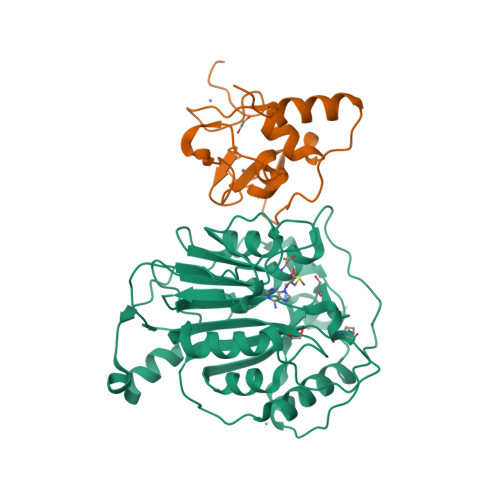
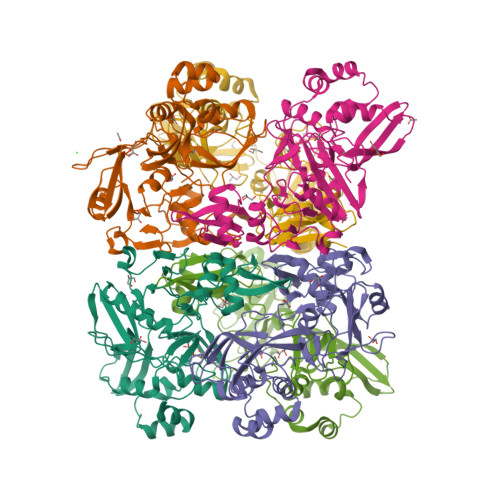
Selected 3D Structures  |
|||||||||||
|
|
||||||||||
|
|
||||||||||
|
|
||||||||||
|
|
||||||||||
| General Comments |
|
The pp1ab polyprotein is a pro-protein, that is proteolytically cleaved to form 16 shorter proteins. The first 11 proteins are also cleaved from pp1a. Nsps 12-16 are unique to pp1ab [18]. Most of what is predicted about the functions of these proteins is implied by similarities to orthologues from other coronaviruses [5-6]. Non-structural protein 12 behaves as a RNA-dependent RNA polymerase (RdRp) that copies the viral RNA template [2]. This is the molecular target of nucleotide analogue antiviral drugs like remdesivir, tenofovir and ribavirin [7,17]. It has a zinc-binding domain in its N-terminus and its activity is magnesium-dependent [1,8,15]. Cryo-EM resolution of the SARS-CoV-2 polymerase complex architecture shows that it contains the nsp12 catalytic subunit and nsp7-nsp8 co-factors [10]. Experimental evidence reveals that nsp12 antagonises the host antiviral type I interferon response to SARS-CoV-2 infection by suppresseing nuclear translocation of IRF3 [16]. Non-structural protein 13 has helicase unwinding activity. Small molecule inhibitors of MERS-CoV helicase have been reported [19]. There is a report of a novel allosteric nsp13 inhibitor that was designed for SARS-CoV-2 antiviral potential [12]. Non-structural protein 14 is a manganese-dependent uridylate-specific endoribonuclease with proofreading exoribonuclease activity that is required for fidelity during viral RNA synthesis. A nsp10/nsp14 complex exerts potent exoribonuclease activity that may be connected to a replicative mismatch repair mechanism [4]. The presence of nsp14 is predicted to be responsible for the high resistance of coronaviruses to many nucleoside analogue inhibitors [11]. Non-structural protein 15 (NendoU) is a uridylate-specific endoribonuclease [20]. The catalytic activity of MERS-CoV nsp15 can be modulated by interactions with other nsps (e.g. nsp8 and a nsp7/8 complex), oligomeric assembly and RNA binding efficiency. Nsp15 is essential in coronavirus biology, but interestingly its deletion does not disrupt replication, so its principal action remains elusive. A resolved X-ray structure has been published and deposited with the RCSB Protein Data Bank (6W01) [9]. This shows that enzymatic functional arises from a hexameric complex whose similarity to SARS-CoV nsp15 suggests that inhibitors of this older coronavirus may also inhibit the more recently identified one. MERS-CoV nsp15 inhibitors are considered to be unlikely to inhibit SARS-CoV-2 nsp15. Non-structural protein 16 dimerises with nsp10 to form a 2'-O-methyltransferase that mediates 2'-O-ribose methylation to the 5'-cap structure of viral mRNAs [3], and this activity helps to improve viral protein translation, to limit host immune detection and inhibit the host's anti-viral interferon response [14]. High-resolution structures of the SARS-CoV-2 nsp16/10 heterodimer have been reported (see for example PDB ID 6W4H) [13]. Such analysis reveals any domains that could be exploited for the development of antiviral inhibitors, which could act to either prevent substrate interaction or block heterodimer formation. |
1. Adedeji AO, Marchand B, Te Velthuis AJ, Snijder EJ, Weiss S, Eoff RL, Singh K, Sarafianos SG. (2012) Mechanism of nucleic acid unwinding by SARS-CoV helicase. PLoS ONE, 7 (5): e36521. [PMID:22615777]
2. Ahn DG, Choi JK, Taylor DR, Oh JW. (2012) Biochemical characterization of a recombinant SARS coronavirus nsp12 RNA-dependent RNA polymerase capable of copying viral RNA templates. Arch Virol, 157 (11): 2095-104. [PMID:22791111]
3. Bouvet M, Debarnot C, Imbert I, Selisko B, Snijder EJ, Canard B, Decroly E. (2010) In vitro reconstitution of SARS-coronavirus mRNA cap methylation. PLoS Pathog, 6 (4): e1000863. [PMID:20421945]
4. Bouvet M, Imbert I, Subissi L, Gluais L, Canard B, Decroly E. (2012) RNA 3'-end mismatch excision by the severe acute respiratory syndrome coronavirus nonstructural protein nsp10/nsp14 exoribonuclease complex. Proc Natl Acad Sci USA, 109 (24): 9372-7. [PMID:22635272]
5. Dong S, Sun J, Mao Z, Wang L, Lu YL, Li J. (2020) A guideline for homology modeling of the proteins from newly discovered betacoronavirus, 2019 novel coronavirus (2019-nCoV). J Med Virol, 92 (9): 1542-1548. [PMID:32181901]
6. Fung TS, Liu DX. (2019) Human Coronavirus: Host-Pathogen Interaction. Annu Rev Microbiol, 73: 529-557. [PMID:31226023]
7. Gordon CJ, Tchesnokov EP, Woolner E, Perry JK, Feng JY, Porter DP, Götte M. (2020) Remdesivir is a direct-acting antiviral that inhibits RNA-dependent RNA polymerase from severe acute respiratory syndrome coronavirus 2 with high potency. J Biol Chem, 295 (20): 6785-6797. [PMID:32284326]
8. Imbert I, Guillemot JC, Bourhis JM, Bussetta C, Coutard B, Egloff MP, Ferron F, Gorbalenya AE, Canard B. (2006) A second, non-canonical RNA-dependent RNA polymerase in SARS coronavirus. EMBO J, 25 (20): 4933-42. [PMID:17024178]
9. Kim Y, Jedrzejczak R, Maltseva NI, Wilamowski M, Endres M, Godzik A, Michalska K, Joachimiak A. (2020) Crystal structure of Nsp15 endoribonuclease NendoU from SARS-CoV-2. Protein Sci, 29 (7): 1596-1605. [PMID:32304108]
10. Peng Q, Peng R, Yuan B, Zhao J, Wang M, Wang X, Sun Y, Fan Z, Qi J, Gao GF et al.. (2020) Structural and biochemical characterization of nsp12-nsp7-nsp8 core polymerase complex from SARS-CoV-2. Cell Reports,. DOI: 10.1016/j.celrep.2020.107774
11. Pruijssers AJ, Denison MR. (2019) Nucleoside analogues for the treatment of coronavirus infections. Curr Opin Virol, 35: 57-62. [PMID:31125806]
12. Ramsey JR, Shelton PMM, Heiss TK, Olinares PDB, Vostal LE, Soileau H, Grasso M, Casebeer SW, Adaniya S, Miller M et al.. (2024) Using a Function-First "Scout Fragment"-Based Approach to Develop Allosteric Covalent Inhibitors of Conformationally Dynamic Helicase Mechanoenzymes. J Am Chem Soc, 146 (1): 62-67. DOI: 10.1101/2023.09.25.559391 [PMID:38134034]
13. Rosas-Lemus M, Minasov G, Shuvalova L, Inniss NL, Kiryukhina O, Brunzelle J, Satchell KJF. (2020) High-resolution structures of the SARS-CoV-2 2′-O-methyltransferase reveal strategies for structure-based inhibitor design. Science Signaling, 13 (651): eabe1202. DOI: 10.1126/scisignal.abe1202
14. Schindewolf C, Lokugamage K, Vu M, Johnson BA, Scharton D, Plante JA, Kalveram BK, Crocquet-Valdes PA, Sotcheff SL, Jaworski E et al.. (2022) SARS-CoV-2 Uses Nonstructural Protein 16 to Evade Restriction by IFIT1 and IFIT3. bioRxiv, Preprint. DOI: 10.1101/2022.09.26.509529
15. Tanner JA, Watt RM, Chai YB, Lu LY, Lin MC, Peiris JS, Poon LL, Kung HF, Huang JD. (2003) The severe acute respiratory syndrome (SARS) coronavirus NTPase/helicase belongs to a distinct class of 5' to 3' viral helicases. J Biol Chem, 278 (41): 39578-82. [PMID:12917423]
16. Wang W, Zhou Z, Xiao X, Tian Z, Dong X, Wang C, Li L, Ren L, Lei X, Xiang Z et al.. (2021) SARS-CoV-2 nsp12 attenuates type I interferon production by inhibiting IRF3 nuclear translocation. Cell Mol Immunol, 18 (4): 945-953. [PMID:33637958]
17. Wang Y, Anirudhan V, Du R, Cui Q, Rong L. (2021) RNA-dependent RNA polymerase of SARS-CoV-2 as a therapeutic target. J Med Virol, 93 (1): 300-310. [PMID:32633831]
18. Yoshimoto FK. (2020) The Proteins of Severe Acute Respiratory Syndrome Coronavirus-2 (SARS CoV-2 or n-COV19), the Cause of COVID-19. The Protein Journal volume, 39: 198–216. DOI: 10.1007/s10930-020-09901-4
19. Zaher NH, Mostafa MI, Altaher AY. (2020) Design, synthesis and molecular docking of novel triazole derivatives as potential CoV helicase inhibitors. Acta Pharm, 70 (2): 145-159. [PMID:31955138]
20. Zhang L, Li L, Yan L, Ming Z, Jia Z, Lou Z, Rao Z. (2018) Structural and Biochemical Characterization of Endoribonuclease Nsp15 Encoded by Middle East Respiratory Syndrome Coronavirus. J Virol, 92 (22). [PMID:30135128]